 Clickpoints
Clickpoints
Click Points is a program written in the Python programming language, which serves on the one hand as an image viewer and on the other hand as an data display and annotation tool. Every frame can be annotated by a description, marked points/tracks, or marked areas (paint brush). This helps to view image data, do manual evaluation of data, help to create semi-automatic evaluation or display the results of automatic image evaluation.
Bayesloop
 bayesloop is a python module that focuses on fitting time series models with time-varying parameters and model selection based on Bayesian inference. Instead of relying on MCMC methods, bayesloop uses a grid-based approach to evaluate probability distributions, allowing for an efficient approximation of the marginal likelihood (evidence). The marginal likelihood represents a powerful tool to objectively compare different models and/or optimize the hyper-parameters of hierarchical models. To avoid the curse of dimensionality when analyzing time series models with time-varying parameters, bayesloop employs a sequential inference algorithm that is based on the forward-backward-algorithm used in Hidden Markov models. Here, the relevant parameter spaces are kept low-dimensional by processing time series data step by step. The module covers a large class of time series models and is easily extensible.
bayesloop is a python module that focuses on fitting time series models with time-varying parameters and model selection based on Bayesian inference. Instead of relying on MCMC methods, bayesloop uses a grid-based approach to evaluate probability distributions, allowing for an efficient approximation of the marginal likelihood (evidence). The marginal likelihood represents a powerful tool to objectively compare different models and/or optimize the hyper-parameters of hierarchical models. To avoid the curse of dimensionality when analyzing time series models with time-varying parameters, bayesloop employs a sequential inference algorithm that is based on the forward-backward-algorithm used in Hidden Markov models. Here, the relevant parameter spaces are kept low-dimensional by processing time series data step by step. The module covers a large class of time series models and is easily extensible.
The underlying algorithm of bayesloop has been successfully employed in cancer research, studying the migration paths of invasive tumor cells, see this article. For a step-by-step introduction to bayesloop, see the documentation.

Magnetic Tweezer
The high force magnetic tweezer device is a scientific rheometer used to study the mechanical properties of biological materials in the micrometer range, especially the cytoskeleton of cells. Precise forces can be applied to micrometer sized beads bound to cells by focal adhesion complexes.
3-D Traction Microscopy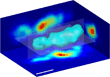
Traction forces in 3-D are important, for instance, for the migration of cells (such as cancer cells, or immune cells) through the connective tissue. To measure forces, we extend ideas from 2-D traction microscopy to the third dimension. In analogy to 2-D traction microscopy, 3-D tractions can be calculated by measuring the 3-D deformation field of the connective tissue matrix surrounding a cell. The image shows the elastic strain energy stored in the extracellular matrix surrounding an invaded breast carcinoma cell.
Cell Stretcher
In order to investigate the mechanical response of cells to uniaxial stretch we use a cell stretching device. Cells are seeded on an flexible PDMS-membrane and are stretched by a stepper motor wich is connected to one side oft the chamber. This is possible in both a two-dimensional and three-dimensional environment.
Biofabrication
Biofabrication by 3-D printing of bioinks requires that the user adjusts the printing pressure such that the resulting flow through the printing needle matches the speed of movement of the printing head. Because most bioinks are highly shear-thinning, small pressure changes result in large changes of the flow speed. Moreover, the user perhaps needs to know the shear stresses inside the printing needle that act upon the cells. The research group of Stephan Gekle (Department of Physics, University of Bayreuth) in collaboration with our group (Richard Gerum) has developed a web-based tool to compute the pressure-flow relationship of shear-thinning bioinks as well as the flow and shear stress profile insde the printing needle.

